Frontiers | Endogenous CRISPR-Cas Systems in Group I Clostridium botulinum and Clostridium sporogenes Do Not Directly Target the Botulinum Neurotoxin Gene Cluster | Microbiology

Introducing Large Genomic Deletions in Human Pluripotent Stem Cells Using CRISPR‐Cas3 - Hou - 2022 - Current Protocols - Wiley Online Library
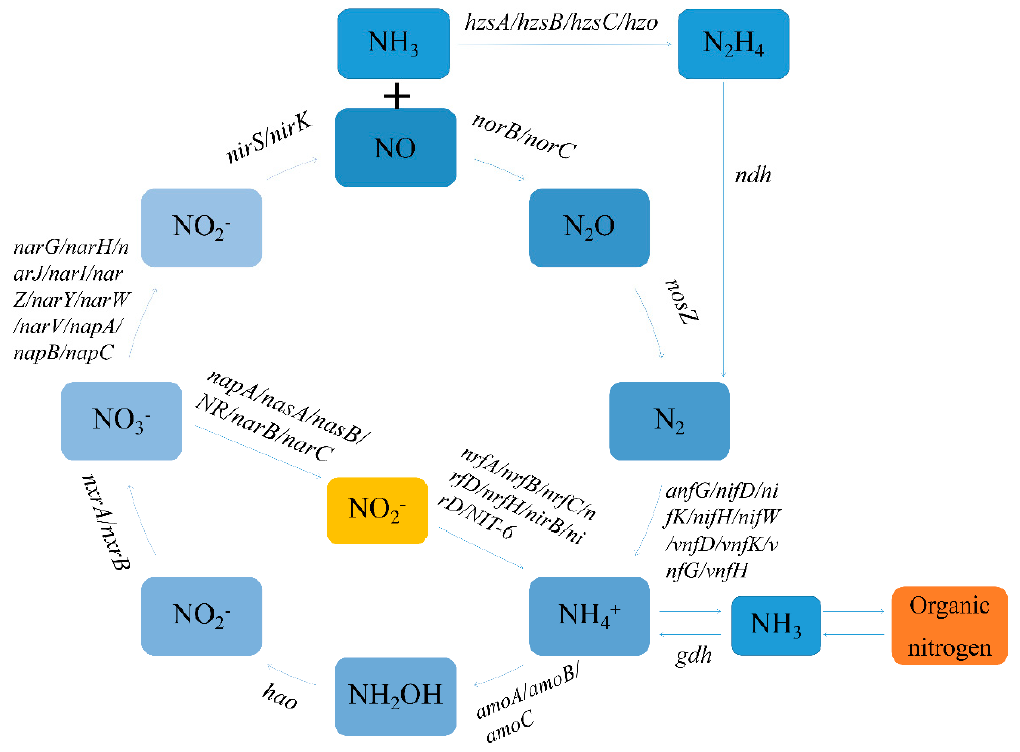
Agriculture | Free Full-Text | Relationship between Plant Roots, Rhizosphere Microorganisms, and Nitrogen and Its Special Focus on Rice | HTML

Efficient genome editing of an extreme thermophile, Thermus thermophilus, using a thermostable Cas9 variant | Scientific Reports
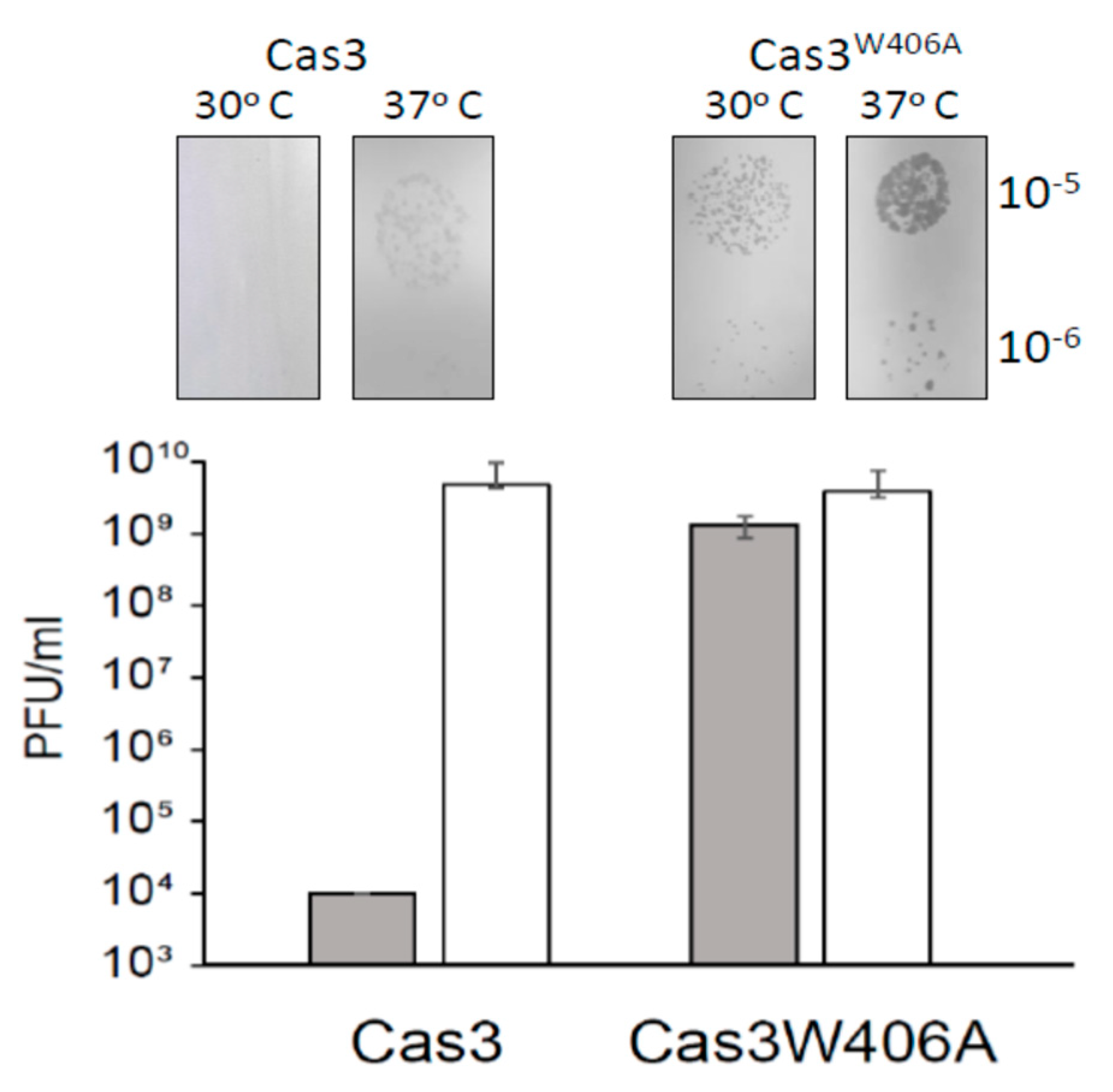
IJMS | Free Full-Text | A Tryptophan 'Gate' in the CRISPR-Cas3 Nuclease Controls ssDNA Entry into the Nuclease Site, That When Removed Results in Nuclease Hyperactivity | HTML
Frontiers | Recent Advanced Technologies for the Characterization of Xenobiotic-Degrading Microorganisms and Microbial Communities | Bioengineering and Biotechnology
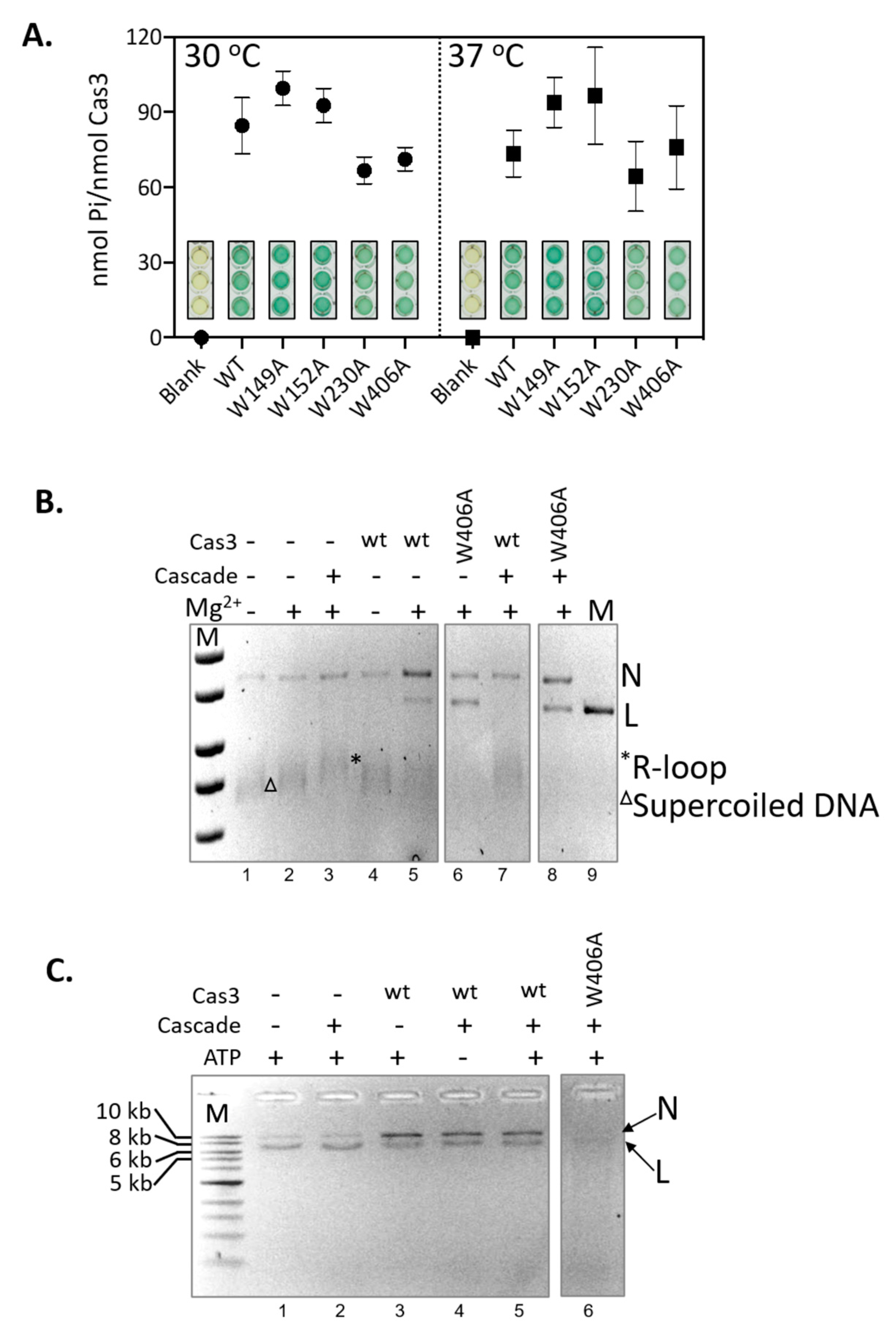
IJMS | Free Full-Text | A Tryptophan 'Gate' in the CRISPR-Cas3 Nuclease Controls ssDNA Entry into the Nuclease Site, That When Removed Results in Nuclease Hyperactivity | HTML